Journal of South China University of Technology(Natural Science Edition) ›› 2026, Vol. 54 ›› Issue (2): 77-90.doi: 10.12141/j.issn.1000-565X.250230
• Computer Science & Technology • Previous Articles Next Articles
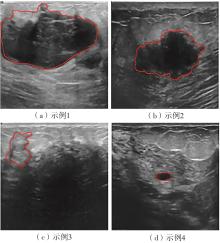
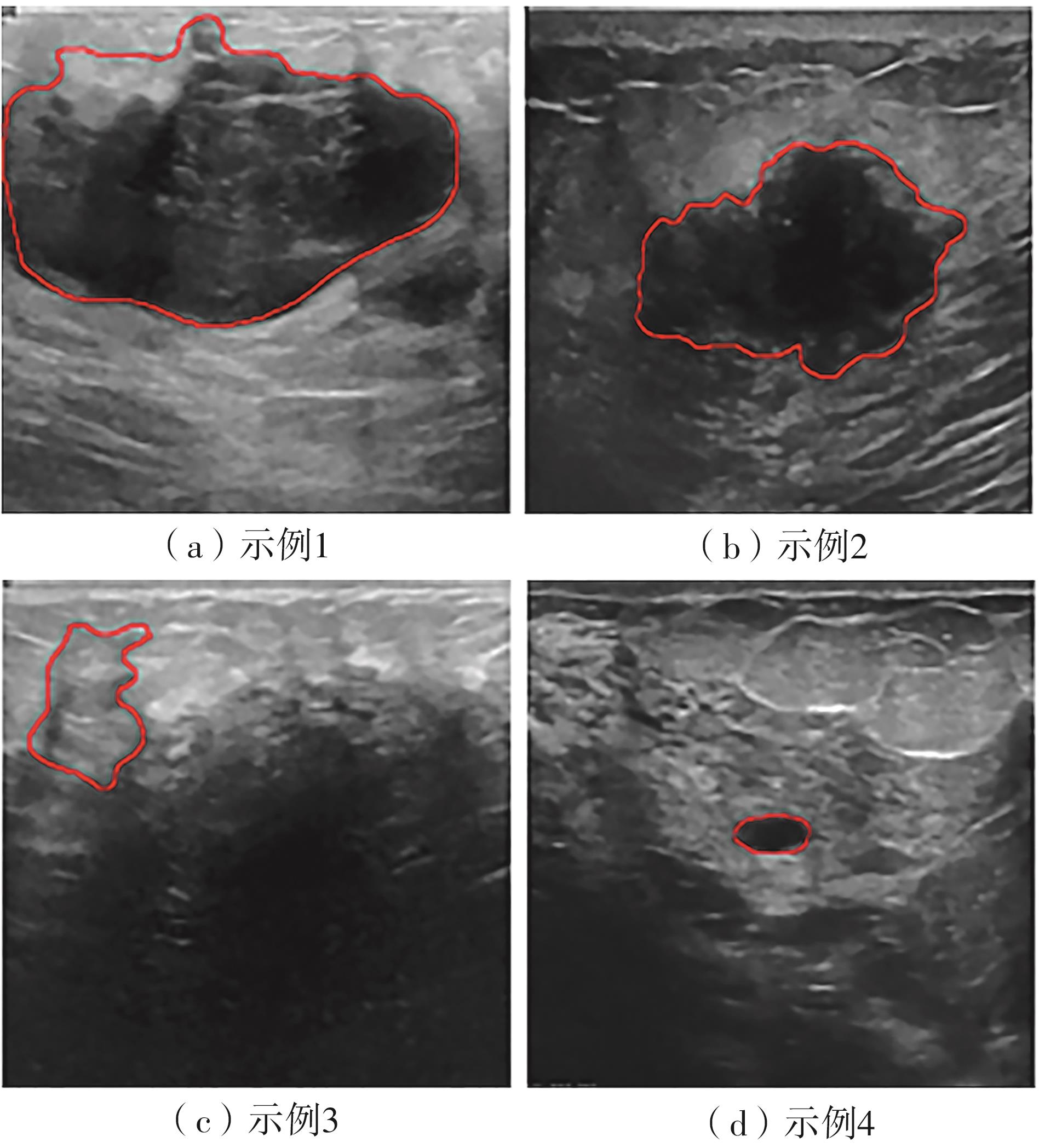
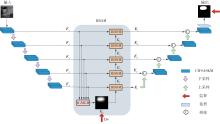
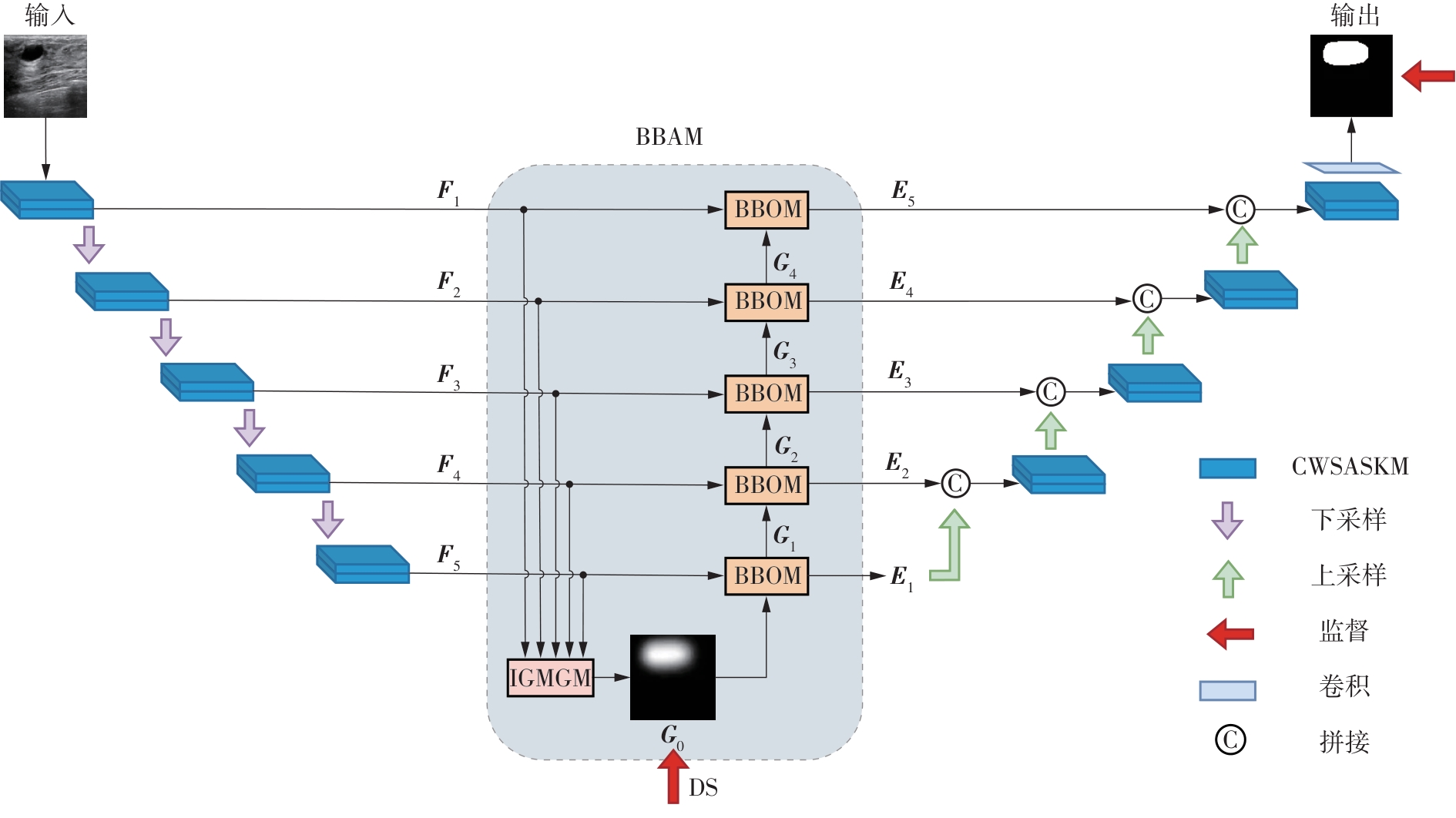
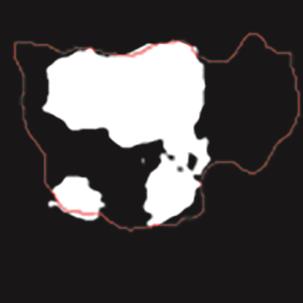
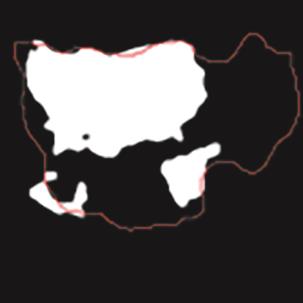
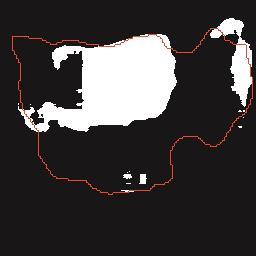
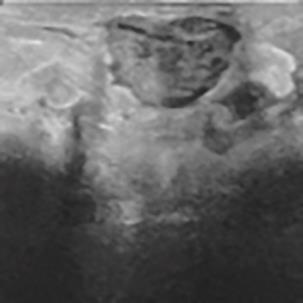
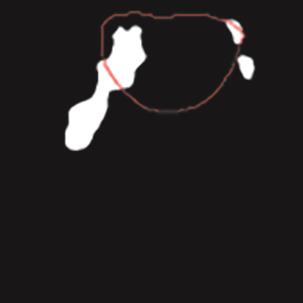
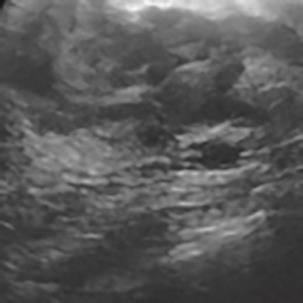
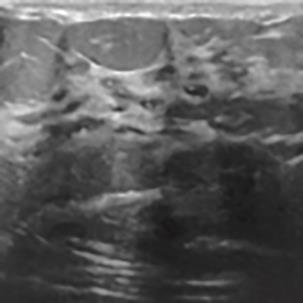
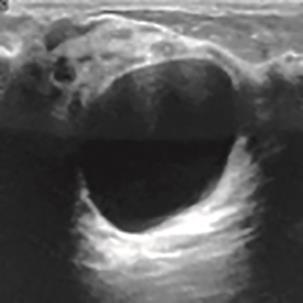
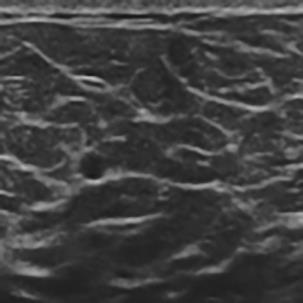
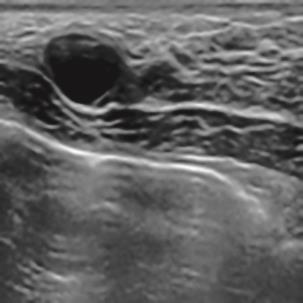
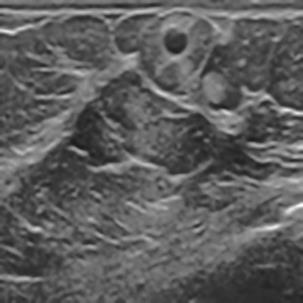
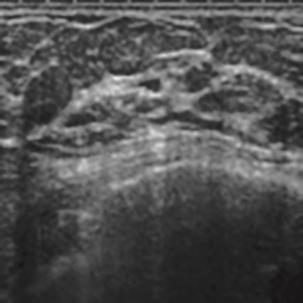
A Breast Ultrasound Images Lesion Segmentation Network Based on Channel-Wise Spatially Adaptive Selective Kernel Convolution and Bidirectional Boundary-Aware Mechanism
WANG Jie, LI Luyao
- School of Computer Science,Beijing University of Technology,Beijing 100124,China
-
Received:2025-07-14Online:2026-02-25Published:2025-09-01 -
About author:王洁(1972—),女,博士,副教授,主要从事逻辑程序设计、面向Agent的程序语言和深度学习研究。E-mail: wj@bjut.edu.cn -
Supported by:the National Natural Science Foundation of China(62476015)
CLC Number:
Cite this article
WANG Jie, LI Luyao. A Breast Ultrasound Images Lesion Segmentation Network Based on Channel-Wise Spatially Adaptive Selective Kernel Convolution and Bidirectional Boundary-Aware Mechanism[J]. Journal of South China University of Technology(Natural Science Edition), 2026, 54(2): 77-90.
share this article
Table 1
Impact of compressed channel numbers on network segmentation performance"
| 数据集 | 通道数 | 参数量/106 | 均值±标准差 | |||
|---|---|---|---|---|---|---|
| J/% | P/% | R/% | cDice/% | |||
| BUSI | 8 | 14.60 | 71.66 ± 2.10 | 82.74 ± 2.67 | 80.89 ± 2.30 | 80.08 ± 2.11 |
| 16 | 14.63 | 71.97 ± 1.84 | 82.85 ± 1.05 | 81.40 ± 2.22 | 80.44 ± 1.73 | |
| 32 | 14.73 | 71.85 ± 1.92 | 83.12 ± 1.68 | 81.25 ± 2.45 | 80.52 ± 1.88 | |
| UDIAT | 8 | 14.60 | 77.82 ± 2.45 | 87.95 ± 4.12 | 86.45 ± 1.78 | 85.88 ± 2.19 |
| 16 | 14.63 | 78.14 ± 2.39 | 88.31 ± 3.99 | 86.73 ± 1.62 | 86.10 ± 2.13 | |
| 32 | 14.73 | 78.35 ± 2.85 | 88.55 ± 4.35 | 86.85 ± 2.10 | 86.35 ± 2.55 | |
Table 2
Comparison of segmentation performance among different methods on BUSI dataset"
| 方法 | 均值±标准差 | |||
|---|---|---|---|---|
| J/% | P/% | R/% | cDice/% | |
| U-Net | 61.31 ± 1.74 | 75.91 ± 1.85 | 73.84 ± 1.87 | 71.57 ± 1.85 |
| UNet++ | 64.66 ± 1.39 | 75.65 ± 2.58 | 78.15 ± 2.17 | 73.67 ± 1.62 |
| Attention U-Net | 64.87 ± 1.89 | 76.88 ± 1.93 | 77.39 ± 4.32 | 73.82 ± 2.22 |
| MADU-Net | 64.69 ± 1.34 | 78.72 ± 1.10 | 75.88 ± 1.82 | 74.29 ± 1.39 |
| STAN | 65.72 ± 2.89 | 80.27 ± 2.59 | 76.13 ± 2.94 | 74.96 ± 2.88 |
| SK U-Net | 69.08 ± 1.99 | 80.88 ± 1.58 | 78.41 ± 1.92 | 77.44 ± 1.77 |
| AE U-Net | 66.81 ± 1.77 | 80.44 ± 2.13 | 76.45 ± 2.37 | 75.64 ± 1.86 |
| AAU-Net | 69.35 ± 2.60 | 80.17 ± 2.45 | 79.87 ± 2.08 | 77.91 ± 2.25 |
| ESKNet | 70.28 ± 1.96 | 81.80 ± 1.18 | 80.04 ± 1.35 | 78.60 ± 1.79 |
| MBDPNet | 69.19 ± 2.38 | 80.41 ± 0.98 | 80.12 ± 2.38 | 78.21 ± 2.09 |
| CWSASKM-BBAM-Net | 71.97 ± 1.84 | 82.85 ± 1.05 | 81.40 ± 2.22 | 80.44 ± 1.73 |
Table 3
Comparison of segmentation performance among different methods on UDIAT dataset"
| 方法 | 均值±标准差 | |||
|---|---|---|---|---|
| J/% | P/% | R/% | cDice/% | |
| U-Net | 70.30 ± 3.37 | 81.53 ± 2.34 | 81.86 ± 3.38 | 79.43 ± 2.71 |
| UNet++ | 74.30 ± 4.17 | 86.27 ± 3.63 | 82.72 ± 4.40 | 82.37 ± 3.59 |
| Attention U-Net | 73.69 ± 3.78 | 84.93 ± 3.95 | 83.25 ± 2.25 | 81.93 ± 2.76 |
| MADU-Net | 71.44 ± 5.91 | 84.03 ± 2.56 | 80.00 ± 5.96 | 79.86 ± 4.55 |
| STAN | 67.76 ± 4.09 | 81.77 ± 3.85 | 77.75 ± 4.45 | 76.49 ± 3.65 |
| SK U-Net | 74.54 ± 3.75 | 84.51 ± 5.69 | 84.32 ± 1.84 | 82.69 ± 3.75 |
| AE U-Net | 72.43 ± 3.33 | 84.22 ± 2.74 | 82.91 ± 3.65 | 80.90 ± 2.62 |
| AAU-Net | 73.76 ± 2.81 | 84.82 ± 3.37 | 83.01 ± 2.93 | 82.20 ± 2.72 |
| ESKNet | 75.39 ± 2.39 | 85.33 ± 3.06 | 86.17 ± 2.01 | 84.09 ± 2.33 |
| MBDPNet | 75.16 ± 2.54 | 84.82 ± 3.27 | 86.09 ± 1.65 | 83.81 ± 2.02 |
| CWSASKM-BBAM-Net | 78.14 ± 2.39 | 88.31 ± 3.99 | 86.73 ± 1.62 | 86.10 ± 2.13 |
Table 6
Validation results of different methods trained on dataset BUSI and evaluated on dataset STU"
| 方法 | 均值±标准差 | |||
|---|---|---|---|---|
| J/% | P/% | R/% | cDice/% | |
| U-Net | 73.57 ± 1.25 | 85.31 ± 2.49 | 83.80 ± 2.36 | 82.50 ± 1.45 |
| UNet++ | 74.75 ± 0.93 | 84.42 ± 2.73 | 87.46 ± 1.40 | 83.97 ± 0.90 |
| Attention U-Net | 72.30 ± 1.29 | 84.77 ± 2.50 | 82.72 ± 2.56 | 81.51 ± 1.55 |
| MADU-Net | 72.65 ± 0.84 | 85.25 ± 1.40 | 83.30 ± 1.54 | 82.59 ± 0.69 |
| STAN | 74.68 ± 0.63 | 84.09 ± 0.84 | 86.73 ± 1.40 | 84.31 ± 0.91 |
| SK U-Net | 77.41 ± 0.39 | 84.08 ± 0.28 | 90.24 ± 0.37 | 86.13 ± 0.30 |
| AE U-Net | 73.78 ± 0.64 | 83.07 ± 0.92 | 86.60 ± 1.43 | 83.65 ± 0.97 |
| AAU-Net | 76.75 ± 1.00 | 83.49 ± 1.01 | 91.49 ± 0.58 | 86.21 ± 0.76 |
| ESKNet | 77.60 ± 1.01 | 84.20 ± 1.09 | 91.71 ± 0.56 | 86.78 ± 0.75 |
| MBDPNet | 78.16 ± 0.51 | 84.79 ± 0.35 | 90.46 ± 0.34 | 86.67 ± 0.34 |
| CWSASKM-BBAM-Net | 78.47 ± 0.90 | 84.84 ± 1.30 | 92.36 ± 0.85 | 87.40 ± 0.64 |
Table 7
Validation results of different methods trained on dataset UDIAT and evaluated on dataset STU"
| 方法 | 均值±标准差 | |||
|---|---|---|---|---|
| J/% | P/% | R/% | cDice/% | |
| U-Net | 70.28 ± 0.83 | 91.98 ± 0.67 | 75.80 ± 1.27 | 80.32 ± 0.82 |
| UNet++ | 72.45 ± 2.69 | 93.23 ± 1.39 | 76.99 ± 3.06 | 81.77 ± 2.55 |
| Attention U-Net | 70.26 ± 2.50 | 91.51 ± 0.84 | 75.38 ± 3.53 | 79.87 ± 2.22 |
| MADU-Net | 68.36 ± 1.84 | 90.75 ± 1.04 | 73.56 ± 2.61 | 78.16 ± 1.90 |
| STAN | 73.74 ± 0.89 | 91.78 ± 0.92 | 79.93 ± 0.68 | 83.35 ± 0.58 |
| SK U-Net | 72.67 ± 1.99 | 92.76 ± 0.97 | 77.58 ± 2.31 | 82.44 ± 1.57 |
| AE U-Net | 71.80 ± 2.05 | 92.85 ± 0.75 | 76.58 ± 2.37 | 81.75 ± 1.63 |
| AAU-Net | 77.36 ± 1.49 | 93.41 ± 0.34 | 82.39 ± 1.49 | 86.28 ± 1.09 |
| ESKNet | 75.81 ± 1.94 | 93.83 ± 0.24 | 80.52 ± 2.17 | 84.58 ± 1.75 |
| MBDPNet | 76.88 ± 1.28 | 93.06 ± 0.40 | 82.20 ± 1.18 | 85.96 ± 1.03 |
| CWSASKM-BBAM-Net | 78.55 ± 1.05 | 94.16 ± 0.45 | 83.25 ± 0.88 | 87.04 ± 0.85 |
Table 8
Quantitative results of ablation experiments on BUSI dataset"
| 模型 | ESK | CWSASKM | BBAM | DS | 均值±标准差 | |||
|---|---|---|---|---|---|---|---|---|
| J/% | P/% | R/% | cDice/% | |||||
| 1 | 61.31 ± 1.74 | 75.91 ± 1.85 | 73.84 ± 1.87 | 71.57 ± 1.85 | ||||
| 2 | √ | 69.06 ± 1.69 | 80.67 ± 1.15 | 79.87 ± 2.60 | 78.27 ± 1.61 | |||
| 3 | √ | 70.19 ± 2.33 | 81.49 ± 1.68 | 80.26 ± 2.25 | 79.03 ± 2.05 | |||
| 4 | √ | 66.19 ± 2.03 | 78.34 ± 1.75 | 76.42 ± 2.01 | 76.68 ± 1.09 | |||
| 5 | √ | √ | 71.05 ± 2.18 | 82.56 ± 1.51 | 80.47 ± 1.97 | 79.54 ± 1.99 | ||
| 6 | √ | √ | √ | 71.97 ± 1.84 | 82.85 ± 1.05 | 81.40 ± 2.22 | 80.44 ± 1.73 | |
Table 9
Quantitative results of ablation experiments on UDIAT dataset"
| 模型 | ESK | CWSASKM | BBAM | DS | 均值±标准差 | |||
|---|---|---|---|---|---|---|---|---|
| J/% | P/% | R/% | cDice/% | |||||
| 1 | 70.30 ± 3.37 | 81.53 ± 2.34 | 81.86 ± 3.38 | 79.43 ± 2.71 | ||||
| 2 | √ | 74.54 ± 2.08 | 85.70 ± 3.14 | 84.48 ± 1.36 | 83.33 ± 2.25 | |||
| 3 | √ | 75.08 ± 2.61 | 84.11 ± 3.49 | 85.37 ± 1.37 | 83.56 ± 2.58 | |||
| 4 | √ | 72.25 ± 2.83 | 82.51 ± 2.51 | 82.71 ± 2.60 | 80.41 ± 2.60 | |||
| 5 | √ | √ | 77.48 ± 2.02 | 87.00 ± 2.97 | 86.25 ± 0.81 | 85.26 ± 1.62 | ||
| 6 | √ | √ | √ | 78.14 ± 2.39 | 88.31 ± 3.99 | 86.73 ± 1.62 | 86.10 ± 2.13 | |
| [1] | BRAY F, LAVERSANNE M, SUNG H,et al .Global cancer statistics 2022:GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries[J].CA:A Cancer Journal for Clinicians,2024,74(3):229-263. |
| [2] | KIM J, HARPER A, McCORMACK V,et al .Global patterns and trends in breast cancer incidence and mortality across 185 countries[J].Nature Medicine,2025,31(4):1154-1162. |
| [3] | FERLAY J, COLOMBET M, SOERJOMATARAM I,et al .Estimating the global cancer incidence and mortality in 2018:GLOBOCAN sources and methods[J].International Journal of Cancer,2019,144(8):1941-1953. |
| [4] | GHONCHEH M, POURNAMDAR Z, SALEHINIYA H .Incidence and mortality and epidemiology of breast cancer in the world[J].Asian Pacific Journal of Cancer Prevention,2016,17(S3):43-46. |
| [5] | JIN J .Breast cancer screening:benefits and harms[J].JAMA,2014,312(23):2585. |
| [6] | MARMOT M G, ALTMAN D G, CAMERON D A,et al .The benefits and harms of breast cancer screening:an independent review[J].British Journal of Cancer,2013,108:2205-2240. |
| [7] | BERG W A, BANDOS A I, MENDELSON E B,et al .Ultrasound as the primary screening test for breast cancer:analysis from ACRIN 6666[J].Journal of the National Cancer Institute,2016,108(4):djv367/1-8. |
| [8] | JOO S, YANG Y S, MOON W K,et al .Computer-aided diagnosis of solid breast nodules:use of an artificial neural network based on multiple sonographic features[J].IEEE Transactions on Medical Imaging,2004,23(10):1292-1300. |
| [9] | MOON W K, LO C M, CHEN R T,et al .Tumor detection in automated breast ultrasound images using quantitative tissue clustering[J].Medical Physics,2014,41(4):042901/1-9. |
| [10] | RONNEBERGER O, FISCHER P, BROX T .U-Net:convolutional networks for biomedical image segmentation[C]∥ Proceedings of the 18th International Conference on Medical Image Computing and Computer-Assisted Intervention.Munich:Springer,2015:234-241. |
| [11] | ZHOU Z, SIDDIQUEE M M R, TAJBAKHSH N,et al .UNet++:redesigning skip connections to exploit multiscale features in image segmentation[J].IEEE Transactions on Medical Imaging,2020,39:1856-1867. |
| [12] | BYRA M, JAROSIK P, SZUBERT A,et al .Breast mass segmentation in ultrasound with selective kernel U-Net convolutional neural network[J].Biomedical Signal Processing and Control,2020,61:102027/1-10. |
| [13] | CHEN G, LI L, DAI Y,et al .AAU-Net:an adaptive attention U-Net for breast lesions segmentation in ultrasound images[J].IEEE Transactions on Medical Imaging,2023,42:1289-1300. |
| [14] | CHEN G, ZHOU L, ZHANG J,et al .ESKNet:an enhanced adaptive selection kernel convolution for ultrasound breast tumors segmentation[J].Expert Systems with Applications,2024,246:123265/1-17. |
| [15] | ABRAHAM N, KHAN N M .A novel focal Tversky loss function with improved attention U-Net for lesion segmentation[C]∥ Proceedings of IEEE the 16th International Symposium on Biomedical Imaging.Venice:IEEE,2019:683-687. |
| [16] | OKTAY O, SCHLEMPER J, FOLGOC L L,et al .Attention U-Net:learning where to look for the pancreas[C]∥ Proceedings of the 1st Conference on Medical Imaging with Deep Learning.Amsterdam:OpenReview,2018:1-10. |
| [17] | YAN Y, LIU Y, WU Y,et al .Accurate segmentation of breast tumors using AE U-Net with HDC model in ultrasound images[J].Biomedical Signal Processing and Control,2022,72:103299/1-7. |
| [18] | AL-DHABYANI W, GOMAA M, KHALED H,et al .Dataset of breast ultrasound images[J].Data in Brief,2020,28:104863/1-5. |
| [19] | YAP M H, GOYAL M, OSMAN F,et al .Breast ultrasound region of interest detection and lesion localisation[J].Artificial Intelligence in Medicine,2020,107:101880/1-8. |
| [20] | ZHUANG Z, LI N, JOSEPH RAJ A N,et al .An RDAU-NET model for lesion segmentation in breast ultrasound images[J].PLoS One,2019,14(8):e0221535/1-23. |
| [21] | SHAREEF B, XIAN M, VAKANSKI A .STAN:small tumor-aware network for breast ultrasound image segmentation[C]∥ Proceedings of IEEE the 17th International Symposium on Biomedical Imaging.Iowa City:IEEE,2020:1-5. |
| [22] | CUI K, TIAN Q, WANG H,et al .An improved framework for breast ultrasound image segmentation with multiple branches depth perception and layer compression residual module[J].Engineering Applications of Artificial Intelligence,2025,146:110265/1-13. |
| [1] | WANG Yaoqi, LU Yaqi, WANG Xiaopeng. A Long-Range Lane Detection Method with Enhanced Spatial Perception [J]. Journal of South China University of Technology(Natural Science Edition), 2026, 54(2): 62-76. |
| [2] | GUO Lihua, LIN Yanyu, CHEN Ke. An Atomic Structure Segmentation Method for High-Noise STEM Image Based on Materials Structural Priors [J]. Journal of South China University of Technology(Natural Science Edition), 2025, 53(11): 27-36. |
| [3] | CHEN Zhong, WANG Aochen, GAO Xinyi, HE Lihui, ZHANG Xianmin. Multi-Object Real-Time Tracking Method Based on Multi-View Near-Infrared Vision [J]. Journal of South China University of Technology(Natural Science Edition), 2025, 53(7): 31-38. |
| [4] | MA Jinlin, JIU Zhiqing, MA Ziping, XIA Mingge, ZHANG Kai, CHENG Yexia, MA Ruishi. A Liver Tumor Image Segmentation Method Based on Multi-Scale Feature Fusion and Reconstruction Convolution [J]. Journal of South China University of Technology(Natural Science Edition), 2025, 53(5): 94-108. |
| [5] | HU Guanghua, DAI Zhigang, WANG Qinghui. Machining Feature Recognition Method of B-Rep Model Based on Graph Neural Network [J]. Journal of South China University of Technology(Natural Science Edition), 2025, 53(5): 20-31. |
| [6] | MA Xiaoliang, GAO Jie, LIU Ying, PEI Qingqi, ZHAO Ruqiang, YANG Bangxing, DENG Congjian. Customer Service Knowledge Recommendation Large Model Construction Driven by Intent Understanding [J]. Journal of South China University of Technology(Natural Science Edition), 2025, 53(3): 40-49. |
| [7] | ZHANG Yan, YAN Yi, WU Hongying, WANG Sitong, WU Yefeng, WANG Nian. Segmentation Method of Barefoot Footprint Based on Multi-Granularity Feature and Region Relationship [J]. Journal of South China University of Technology(Natural Science Edition), 2025, 53(3): 57-67. |
| [8] | XIAN Jin, XU Xiaoru, XIAN Yunting, XIAN Chuhua. Image Inpainting Algorithm Based on Hybrid Encoding and Mask Space Modulation [J]. Journal of South China University of Technology(Natural Science Edition), 2025, 53(3): 31-39. |
| [9] | LIU Ning, HUA Tianbiao, WANG Gao, CHEN Faming. A Batch Scheduling Method of Flexible Job-Shop with Partially Out-of-Ordered Execute Operation [J]. Journal of South China University of Technology(Natural Science Edition), 2024, 52(10): 51-63. |
| [10] | HU Guanghua, TU Qianxi. Surface Defect Detection Method for Industrial Products Based on Photometric Stereo and Dual Stream Feature Fusion Network [J]. Journal of South China University of Technology(Natural Science Edition), 2024, 52(10): 112-123. |
| [11] | MA Biyun, FAN Yihua, LIU Jiaojiao. Blood Flow Velocity Estimation Method Based on Multi-Frequency Pulse Sampling [J]. Journal of South China University of Technology(Natural Science Edition), 2024, 52(10): 22-30. |
| [12] | YUAN Xixi, CAI Zhanchuan, SHI Wuzhen, YIN Wennan. Image Compression Method Based on the Integer U Transform Algorithm [J]. Journal of South China University of Technology(Natural Science Edition), 2024, 52(10): 124-134. |
| [13] | HU Guanghua, OU Meitong, LI Zhendong. Multi-Object Recognition and 6-DoF Pose Estimation Based on Synthetic Datasets [J]. Journal of South China University of Technology(Natural Science Edition), 2024, 52(4): 42-50. |
| [14] | LUO Yutao, GAO Qiang. Traffic Sign Detection Based on Channel Attention and Feature Enhancement [J]. Journal of South China University of Technology(Natural Science Edition), 2023, 51(12): 64-72. |
| [15] | LI Haiyan, YIN Haolin, LI Peng, et al.. Image Inpainting Algorithm Based on Dense Feature Reasoning and Mix Loss Function [J]. Journal of South China University of Technology(Natural Science Edition), 2023, 51(9): 99-109. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||