Journal of South China University of Technology(Natural Science Edition) ›› 2025, Vol. 53 ›› Issue (5): 94-108.doi: 10.12141/j.issn.1000-565X.240439
• Computer Science & Technology • Previous Articles Next Articles
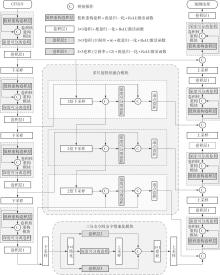
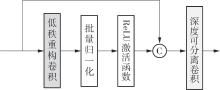
A Liver Tumor Image Segmentation Method Based on Multi-Scale Feature Fusion and Reconstruction Convolution
MA Jinlin1,2, JIU Zhiqing2, MA Ziping3, XIA Mingge2, ZHANG Kai2, CHENG Yexia4, MA Ruishi2
- 1.Key Laboratory of Intelligent Information Processing of Image and Graphics,North Minzu University,Yinchuan 750021,Ningxia,China
2.School of Computer Science and Engineering,North Minzu University,Yinchuan 750021,Ningxia,China
3.School of Mathematics and Information Science,North Minzu University,Yinchuan 750021,Ningxia,China
4.China Mobile Communications Corporation,Beijing 100033,China
-
Received:2024-09-02Online:2025-05-25Published:2024-12-02 -
Contact:酒志青(2000—),女,硕士生,主要从事图像识别研究。 E-mail:2313795280@qq.com -
About author:马金林(1976—),男,博士,副教授,主要从事计算机视觉、图像识别、计算机图形学等研究。E-mail: 624160@qq.com -
Supported by:the National Natural Science Foundation of China(62462001);the Natural Science Foundation of Ningxia Hui Autonomous Region(2024AAC03147)
CLC Number:
Cite this article
MA Jinlin, JIU Zhiqing, MA Ziping, XIA Mingge, ZHANG Kai, CHENG Yexia, MA Ruishi. A Liver Tumor Image Segmentation Method Based on Multi-Scale Feature Fusion and Reconstruction Convolution[J]. Journal of South China University of Technology(Natural Science Edition), 2025, 53(5): 94-108.
share this article
Table 1
Performance comparison of models with different placement positions of low-rank reconstructive convolution (with a rank value of 8)"
实验 序号 | LRC使用位置和层级 | Dice系数/% | IoU值/% | 浮点运算 次数/1010 | 参数量/107 | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 编码器 | 瓶颈层 | 解码器 | |||||||||||
| Ⅰ | Ⅱ | Ⅲ | Ⅳ | Ⅰ | Ⅱ | Ⅲ | Ⅳ | ||||||
| 1 | — | — | — | — | — | — | — | — | — | 94.95 | 90.48 | 5.477 918 | 3.104 |
| 2 | a | a | a | a | 95.48 | 91.42 | 5.104 206 | 2.949 | |||||
| 3 | b | b | b | b | 95.31 | 91.08 | 4.511 551 | 2.791 | |||||
| 4 | b | b | b | b | b | 95.37 | 91.19 | 4.269 959 | 1.847 | ||||
| 5 | a | a | a | a | a | a | a | a | 95.40 | 91.22 | 3.171 471 | 2.322 | |
| 6 | b | b | b | b | b | b | b | b | 95.11 | 90.70 | 3.545 183 | 2.477 | |
| 7 | a+b | a+b | a+b | a+b | 95.20 | 90.91 | 2.578 815 | 2.164 | |||||
| 8 | a+b | 95.35 | 91.16 | 5.115 530 | 1.688 | ||||||||
| 9 | b | a | b | a | b | a | b | a | b | 95.04 | 90.58 | 3.061 999 | 1.408 |
| 10 | a+b | a+b | a+b | a+b | 95.24 | 90.93 | 3.303 591 | 1.976 | |||||
Table 2
Influence of rank value on model performance"
| 实验序号 | 不同rank值下的Dice系数/% | 不同rank值下的IoU值/% | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 2 | 4 | 8 | 16 | 32 | 2 | 4 | 8 | 16 | 32 | |||
| 2 | 95.03 | 95.22 | 95.48 | 95.38 | 95.11 | 90.58 | 90.91 | 91.42 | 91.25 | 90.78 | ||
| 3 | 95.01 | 95.16 | 95.31 | 95.09 | 95.08 | 90.54 | 90.82 | 91.08 | 90.68 | 90.67 | ||
| 4 | 95.12 | 95.21 | 95.37 | 95.27 | 94.96 | 90.71 | 90.88 | 91.19 | 91.03 | 90.47 | ||
| 5 | 94.96 | 95.10 | 95.40 | 95.13 | 95.14 | 90.52 | 90.73 | 91.22 | 90.74 | 90.79 | ||
| 6 | 94.36 | 94.74 | 95.11 | 94.85 | 94.73 | 89.35 | 90.03 | 90.70 | 90.25 | 90.02 | ||
| 8 | 95.14 | 95.07 | 95.35 | 95.32 | 95.39 | 90.79 | 90.86 | 91.16 | 91.10 | 91.24 | ||
Table 3
Results of ablation experiment"
| 消融实验方案 | Dice系数/% | IoU值/% | 浮点运算次数/1010 | 参数量/107 | ||
|---|---|---|---|---|---|---|
| MSFF | TSPP | KRS | ||||
| — | — | — | 94.95 | 90.48 | 5.477 918 | 3.104 |
| √ | 96.66 | 93.54 | 7.079 019 | 3.806 | ||
| √ | 96.24 | 92.77 | 5.437 719 | 3.000 | ||
| √ | 96.85 | 93.90 | 3.629 279 | 2.466 | ||
| √ | √ | 97.16 | 94.50 | 7.038 820 | 3.703 | |
| √ | √ | 97.11 | 94.38 | 3.962 792 | 2.517 | |
| √ | √ | 97.31 | 94.78 | 5.230 380 | 3.168 | |
| √ | √ | √ | 97.56 | 95.25 | 5.190 181 | 3.064 |
Table 4
Performance comparison of the proposed method with mainstream methods on LiTS2017 dataset"
| 方法 | 肝脏图像分割性能 | 肝肿瘤图像分割性能 | ||
|---|---|---|---|---|
| Dice系数/% | IoU值/% | Dice系数/% | IoU值/% | |
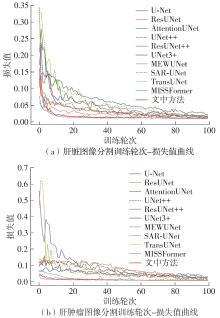
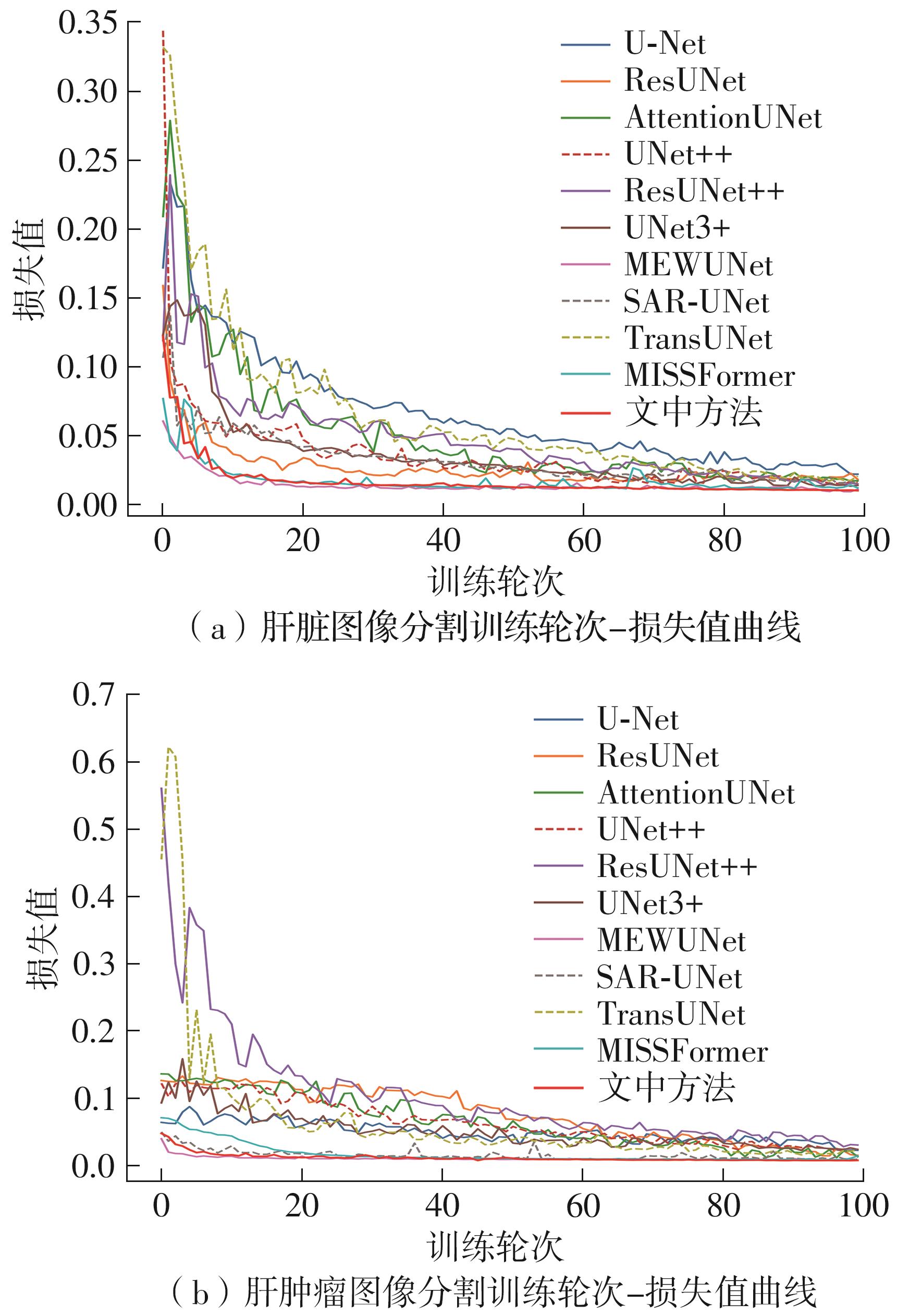
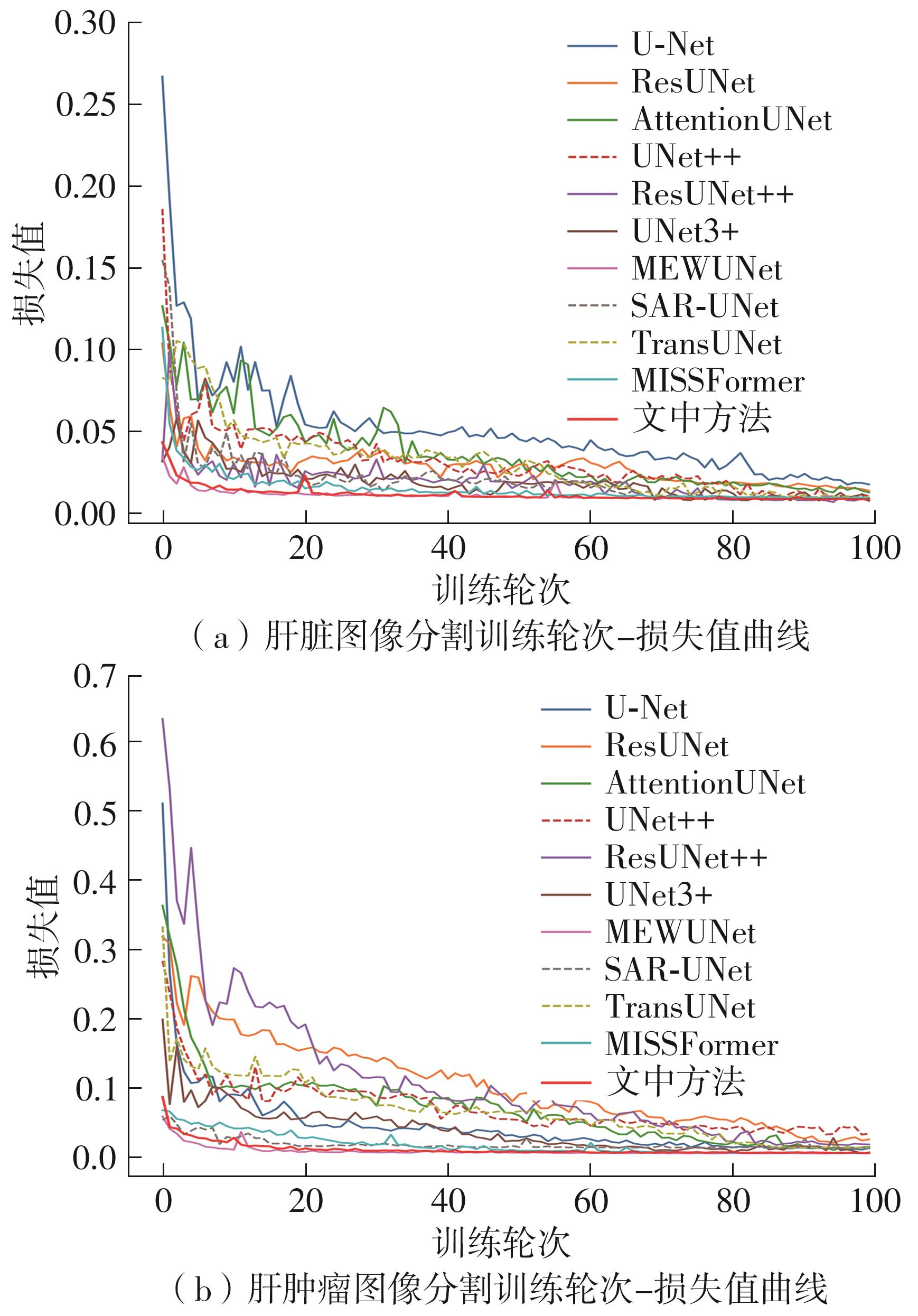
| U-Net[ | 94.95 | 90.48 | 70.88 | 59.51 |
| ResUNet[ | 95.54 | 91.50 | 79.61 | 67.30 |
| AttentionUNet[ | 95.96 | 92.25 | 81.35 | 69.60 |
| UNet++[ | 96.49 | 93.24 | 76.96 | 66.79 |
| ResUNet++[ | 96.58 | 93.42 | 82.07 | 71.13 |
| UNet3+[ | 96.75 | 93.76 | 78.44 | 68.62 |
| MEWUNet[ | 97.28 | 94.74 | 88.26 | 79.66 |
| SAR-UNet[ | 96.39 | 93.18 | 87.23 | 77.89 |
| TransUNet[ | 96.87 | 94.13 | 82.77 | 70.91 |
| MISSFormer[ | 97.32 | 94.77 | 89.43 | 81.02 |
| 文中方法 | 97.56 | 95.25 | 89.71 | 81.58 |
Table 5
Performance comparison of the proposed method with mainstream methods on 3DIRCADb dataset"
| 方法 | 肝脏图像分割性能 | 肝肿瘤图像分割性能 | ||
|---|---|---|---|---|
| Dice系数/% | IoU值/% | Dice系数/% | IoU值/% | |
| U-Net[ | 95.46 | 91.63 | 72.55 | 58.41 |
| ResUNet[ | 96.17 | 92.72 | 80.13 | 68.36 |
| AttentionUNet[ | 96.38 | 93.13 | 81.17 | 70.54 |
| UNet++[ | 96.78 | 93.85 | 82.11 | 72.30 |
| ResUNet++[ | 96.69 | 93.68 | 82.31 | 71.89 |
| UNet3+[ | 96.88 | 94.03 | 81.14 | 69.21 |
| MEWUNet[ | 97.23 | 94.67 | 89.52 | 81.50 |
| SAR-UNet[ | 96.86 | 94.03 | 87.42 | 78.35 |
| TransUNet[ | 97.04 | 94.32 | 82.98 | 72.07 |
| MISSFormer[ | 97.13 | 94.45 | 89.24 | 80.91 |
| 文中方法 | 97.63 | 95.39 | 89.62 | 81.63 |
Table 6
Comparison of computational and parameter complexity between of the proposed method and mainstream methods"
| 方法 | 浮点运算次数/109 | 参数量/107 |
|---|---|---|
| U-Net[ | 54.779 18 | 3.104 |
| ResUNet[ | 56.431 21 | 3.156 |
| AttentionUNet[ | 66.631 85 | 3.488 |
| UNet++[ | 138.660 54 | 3.663 |
| ResUNet++[ | 70.933 84 | 1.448 |
| UNet3+[ | 199.742 46 | 2.697 |
| MEWUNet[ | 40.520 58 | 13.889 |
| SAR-UNet[ | 86.790 94 | 4.218 |
| TransUNet[ | 24.677 27 | 9.323 |
| MISSFormer[ | 7.254 04 | 3.545 |
| 文中方法 | 51.901 81 | 3.064 |
| 1 | RONNEBERGER O, FISCHER P, BROX T .U-Net:convolutional networks for biomedical image segmentation[C]∥Proceedings of the 18th International Conference on Medical Image Computing and Computer-Assisted Intervention—MICCAI 2015.Munich:Springer International Publishing,2015:234-241. |
| 2 | LV P, WANG J, ZHANG X,et al .Deep supervision and atrous inception-based U-Net combining CRF for automatic liver segmentation from CT[J].Scientific Reports,2022,12(1):16995/1-14. |
| 3 | GUO C, SZEMENYEI M, YI Y,et al .SA-UNet:spatial attention U-Net for retinal vessel segmentation[C]∥Proceedings of 2020 25th International Conference on Pattern Recognition.Milan:IEEE,2021:1236-1242. |
| 4 | CAI Z, XIN J, SHI P,et al .DSTUNet: UNet with efficient dense SWIN transformer pathway for medical image segmentation[C]∥Proceedings of 2022 IEEE 19th International Symposium on Biomedical Imaging.Kolkata:IEEE,2022:1-5. |
| 5 | KIRAN I, RAZA B, IJAZ A,et al .DenseRes-UNet: segmentation of overlapped/clustered nuclei from multi organ histopathology images[J].Computers in Bio-logy and Medicine,2022,143:105267/1-15. |
| 6 | ANSARI M Y, YANG Y, BALAKRISHNAN S,et al .A lightweight neural network with multiscale feature enhancement for liver CT segmentation[J].Scientific reports,2022,12(1):14153/1-12. |
| 7 | ÖZCAN F, UÇAN O N, KARAÇAM S,et al .Fully automatic liver and tumor segmentation from CT image using an AIM-Unet[J].Bioengineering,2023,10(2):215/1-21. |
| 8 | AMER A, LAMBROU T, YE X .MDA-Unet: a multi-scale dilated attention U-Net for medical image segmentation[J].Applied Sciences,2022,12(7):3676/1-18. |
| 9 | IBTEHAZ N, KIHARA D .ACC-UNet: a completely convolutional UNet model for the 2020s[C]∥Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention.Cham:Springer Nature Switzerland,2023:692-702. |
| 10 | KONAR D, BHATTACHARYYA S, GANDHI T K,et al .3-D quantum-inspired self-supervised tensor network for volumetric segmentation of medical images[J].IEEE Transactions on Neural Networks and Learning Systems,2024,35(8):10312-10325. |
| 11 | SELVAN R, DAM E B, PETERSEN J .Segmenting two-dimensional structures with strided tensor networks[C]∥Proceedings of the 27th International Conference on Information Processing in Medical ImagingVirtual Event:Springer International Publishing,2021:401-414. |
| 12 | DENTON E L, ZAREMBA W, BRUNA J,et al .Exploiting linear structure within convolutional networks for efficient evaluation[C]∥Proceedings of the Confe-rence on Advances in Neural Information Processing Systems.Montreal:MIT Press,2014:1269-1277. |
| 13 | CHENG Z, LI B, FAN Y,et al .A novel rank selection scheme in tensor ring decomposition based on reinforcement learning for deep neural networks[C]∥Proceedings of 2020 IEEE International Conference on Acoustics,Speech and Signal Processing (ICASSP).Barcelona:IEEE,2020:3292-3296. |
| 14 | LIU Y, NG M K .Deep neural network compression by tucker decomposition with nonlinear response[J].Knowledge-Based Systems,2022,241:108171/1-12. |
| 15 | FALASCHETTI L, MANONI L, TURCHETTI C .A low-rank CNN architecture for real-time semantic segmentation in visual slam applications[J].IEEE Open Journal of Circuits and Systems,2022,3:115-133. |
| 16 | WANG J, LI S, YU L,et al .SDPN: a slight dual-path network with local-global attention guided for medical image segmentation[J].IEEE Journal of Biomedical and Health Informatics,2023,27(6):2956-2967. |
| 17 | CHEN W, ZHU X, SUN R,et al .Tensor low-rank reconstruction for semantic segmentation[C]∥Proceedings of the 16th European Conference on Computer Vision (ECCV 2020).Glasgow:Springer International Publishing,2020:52-69. |
| 18 | FAN X, LU Y, HOU J,et al .DMC-UNet-based segmentation of lung nodules[J].IEEE Access,2023,11:3322437/1-18. |
| 19 | JHA D, SMEDSRUD P H, RIEGLER M A,et al .RseUNet++: an advanced architecture for medical ima-ge segmentation[C]∥Proceedings of 2019 IEEE International Symposium on Multi-Media.San Diego:IEEE,2019:225-2255. |
| 20 | WANG J, LV P, WANG H,et al .SAR-UNet:squeeze-and-excitation block and atrous spatial pyramid pooling based residual U-Net for automatic liver segmentation in computed tomography[J].Computer Me-thods and Programs in Biomedicine,2021,208:106268/1-16. |
| 21 | FAN T, WANG G, LI Y,et al .MA-Net: a multi-scale attention network for liver and tumor segmentation[J].IEEE Access,2020,8:179656-179665. |
| 22 | LI X, CHEN H, QI X,et al .H-DenseUNet:hybrid densely connected UNet for liver and tumor segmentation from CT volumes[J].IEEE transactions on Medical Imaging,2018,37(12):2663-2674. |
| 23 | SHAO Y, ZHOU K, ZHANG L .CSSNet: cascaded spatial shift network for multi-organ segmentation[J].Computers in Biology and Medicine,2024,170:107955/1-14. |
| 24 | LIANG B, TANG C, ZHANG W,et al .N-Net: an UNet architecture with dual encoder for medical image segmentation[J].Signal,Image and Video Processing,2023,17(6):3073-3081. |
| 25 | XIE J, ZHU R, WU Z,et al .FFUNet: a novel feature fusion makes strong decoder for medical image segmentation[J].IET Signal Processing,2022,16(5):501-514. |
| 26 | XIAO X, LIAN S, LUO Z,et al .Weighted ResUNet for high-quality retina vessel segmentation[C]∥Proceedings of 2018 9th International Conference on Information Technology in Medicine and Education.Hangzhou:IEEE,2018:327-331. |
| 27 | OKTAY O, SCHLEMPER J, FOLGOC L L,et al .AttentionUNet: learning where to look for the pancreas[EB/OL].(2018-04-11)[2024-08-30].. |
| 28 | ZHOU Z, RAHMAN SIDDIQUEE M M, TAJBAKHSH N,et al .UNet++: a nested U-Net architecture for medical image segmentation[C]∥Proceedings of the 4th International Workshop on Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support.Granada:Springer International Publishing,2018:3-11. |
| 29 | HUANG H, LIN L, TONG R,et al .UNet3+: a full-scale connected UNet for medical image segmentation[C]∥Proceedings of 2020 IEEE International Conference on Acoustics,Speech and Signal Processing.Barcelona:IEEE,2020:1055-1059. |
| 30 | RUAN J, XIE M, XIANG S,et al .MEWUNet: multi-axis representation learning in frequency domain for medical image segmentation[EB/OL].(2022-10-25)[2024-08-30].. |
| 31 | CHEN J, LU Y, YU Q,et al .TransUNet: transfor-mers make strong encoders for medical image segmentation[EB/OL].(2021-02-08)[2024-08-30].. |
| 32 | HUANG X, DENG Z, LI D,et al .MISSFormer: an effective transformer for 2d medical image segmentation[J].IEEE Transactions on Medical Imaging,2022,42(5):1484-1494. |
| [1] | LIU Jianguo, FENG Yunjian, JI Guo, et al. Improved Stereo Matching Algorithm Based on PSMNet [J]. Journal of South China University of Technology (Natural Science Edition), 2020, 48(1): 60-69,83. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||