

Effects of Trigger Transposable Element-Derived 1 on the Growth of Hepatocellular Carcinoma
Received date: 2023-04-18
Online published: 2023-06-20
Supported by
the National Key R&D Program of China(2018YFA0800303)
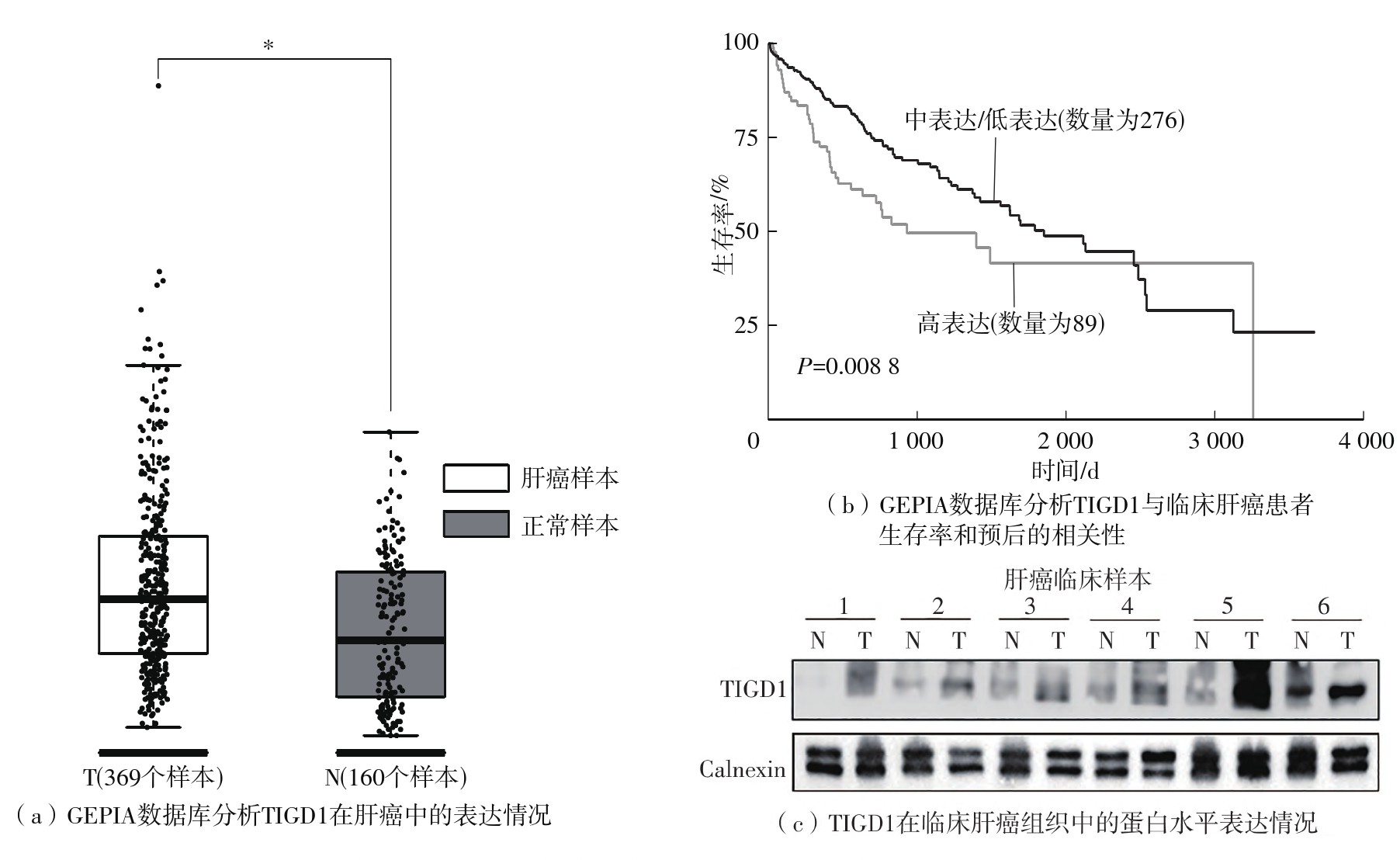
Cell cycle dysregulation is one of the most important hallmarks of cancer. Bioinformatics studies have suggested that trigger transposable element-derived 1 (TIGD1) is expressed at higher levels in the tumor tissues from the clinical hepatocellular carcinoma (HCC) samples and may be related to the cell cycle. However, the underlying mechanism is unclear. To explore the specific mechanism of TIGD1 regulating the growth of hepatocellular carcinoma cells, this paper first analyzed the growth of HCC cell line Hep3B with TIGD1 knockdown by using shRNA plasmid. The results show that cell growth is inhibited. Then, cell cycle analysis by flow cytometry was used to investigate the effect of TIGD1 knockdown on cell cycle of HCC. The results show that the cell cycle progression of the Hep3B cell line is mainly blocked in the G2/M phase. Next, Immunoprecipitation (IP) experiments were used to verify the protein molecules with which TIGD1 might interact. And the results show that TIGD1 may be bound to Aurora kinase interacting protein 1 (AURKAIP1). Further, the Co-IP experiment confirmed the interaction between TIGD1 and AURKAIP1. AURKAIP1 is known to regulate the proteasomal degradation pathway of Aurora kinase A (AURKA), and AURKA is a mitotic regulatory protein that is closely associated with cell cycle progression. The paper further explored the effect of TIGD1 on AURKA protein levels, and the results show that TIGD1 knockdown obviously decreases the protein level of AURKA without affecting its mRNA level in Hep3B cells. In conclusion, TIGD 1 may affect cell cycle progression by regulating the post-transcriptional levels of AURKA in HCC cells, thus affecting the development of HCC.
GAO Ping, YE Zijian, QIAN Xiaoyu, et al . Effects of Trigger Transposable Element-Derived 1 on the Growth of Hepatocellular Carcinoma[J]. Journal of South China University of Technology(Natural Science), 2024 , 52(4) : 1 -7 . DOI: 10.12141/j.issn.1000-565X.230248
| 1 | BRAY F, FERLAY J, SOERJOMATARAM I,et al .Global cancer statistics 2018:GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries[J].CA:A Cancer Journal for Clinicians,2018,68(6):394-424. |
| 2 | VERSLYPE C, Van CUTSEM E, DICATO M,et al .The management of hepatocellular carcinoma.Current expert opinion and recommendations derived from the 10th World Congress on Gastrointestinal Cancer,Barcelona,2008[J].Annals of Oncology,2009,20():vii1-vii6. |
| 3 | HOU W, LIU J, CHEN P,et al .Mutation analysis of key genes in RAS/RAF and PI3K/PTEN pathways in Chinese patients with hepatocellular carcinoma[J].Oncology Letters,2014,8(3):1249-1254. |
| 4 | HUANG M, CHEN C, GENG J,et al. Targeting KDM 1 A attenuates Wnt/β-catenin signaling pathway to eliminate sorafenib-resistant stem-like cells in hepatocellular carcinoma[J].Cancer Letters,2017,398:12-21. |
| 5 | JINDAL A, THADI A, SHAILUBHAI K .Hepatocellular carcinoma:etiology and current and future drugs [J].Journal of Clinical and Experimental Hepatology,2019,9(2):221-232. |
| 6 | LLOVET J M, RICCI S, MAZZAFERRO V,et al .Sorafenib in advanced liver cancer[J].The New England Journal of Medicine,2008,359(4):378-390. |
| 7 | WILLIAMS G H, STOEBER K .The cell cycle and cancer[J].Journal of Pathology,2012,226(2):352-364. |
| 8 | NARASIMHA A M, KAULICH M, SHAPIRO G S,et al .Cyclin D activates the Rb tumor suppressor by mono-phosphorylation[J].eLife,2014,3:e02872/1-21. |
| 9 | TOPACIO B R, ZATULOVSKIY E, CRISTEA S,et al .Cyclin D-Cdk4,6 drives cell-cycle progression via the retinoblastoma protein’s C-terminal helix[J].Molecular Cell,2019,74(4):758-770. |
| 10 | WANG Y, SUN H, WANG Z,et al .Aurora-A:a potential DNA repair modulator[J].Tumour Biology,2014,35(4):2831-2836. |
| 11 | DANG C V .MYC on the path to cancer[J].Cell,2012,149(1):22-35. |
| 12 | SUR S, AGRAWAL D K .Phosphatases and kinases regulating CDC25 activity in the cell cycle:clinical implications of CDC25 overexpression and potential treatment strategies[J].Molecular and Cellular Biochemistry,2016,416(1/2):33-46. |
| 13 | YIN L, YAN J, WANG Y,et al .TIGD1,a gene of unknown function,involves cell-cycle progression and correlates with poor prognosis in human cancer[J].Journal of Cellular Biochemistry,2019,120(6):9758-9767. |
| 14 | GE X, LIU Z, JIAO X,et al .Establishment and validation of a gene signature-based prognostic model to improve survival prediction in adrenocortical carcinoma patients[J].International Journal of Endocrinology,2021,2021:2077633. |
| 15 | FANG G, FAN J, DING Z,et al .Prognostic and predictive value of transcription factors panel for digestive system carcinoma[J].Frontiers in Oncology,2021,11:670129/1-14. |
| 16 | SAJADI M, FAZILTI M, NAZEM H,et al .The expression changes of transcription factors including ANKZF1,LEF1,CASZ1,and ATOH1 as a predictor of survival rate in colorectal cancer:a large-scale analysis[J].Cancer Cell International,2022,22:339/1-9. |
| 17 | ZOU J, ZHANG H, WU Z,et al .TIGD1 is an independent prognostic factor that promotes the progression of colon cancer[J/OL].Cancer Biotherapy and Radiopharmaceuticals,(2022-12-12)[2023-04-15].. |
| 18 | LIM S K, GOPALAN G .Aurora-A kinase interacting protein 1 (AURKAIP1) promotes Aurora-A degradation through an alternative ubiquitin-independent pathway[J].Biochemical Journal,2007,403(1):119-127. |
| 19 | LIM S K, GOPALAN G .Antizyme1 mediates AURKAIP1-dependent degradation of Aurora-A[J].Oncogene,2007,26:6593-6603. |
/
| 〈 |
|
〉 |