

Journal of South China University of Technology(Natural Science) >
Proteomic Analysis of CHO-DG44 Cell Line Expressing Foreign Protein
Received date: 2022-01-18
Online published: 2022-05-12
Supported by
the Marine Economy Development (Six Marine Industries) Special Project of Guangdong Province([GDNRC][2021]50)
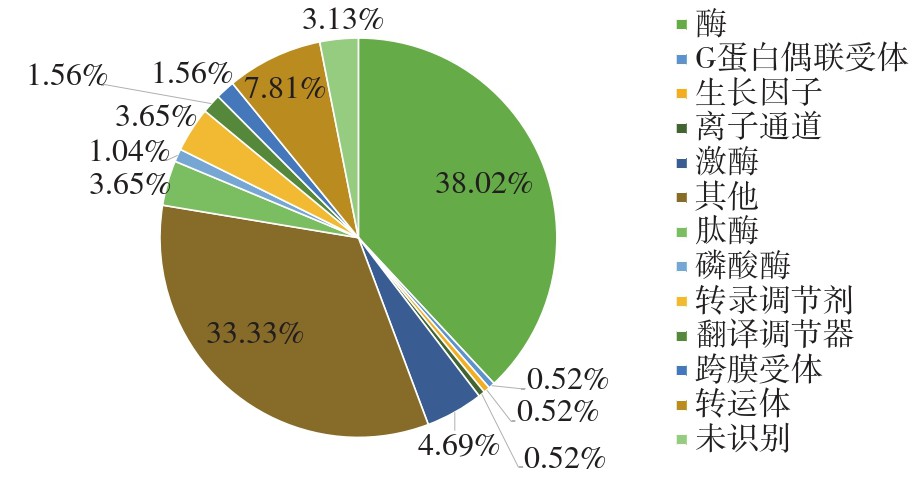
To provide the background for the genetic engineering modification of cell line, this paper compared two CHO-DG44 cells from different sources in terms of their differences in growth, metabolism and expression of exogenous protein TNFR-FC. ITRAQ combined with 2D LC-MS/MS were used to compare the protein expression profiles between CHO-DG44(A)/TNFR-FC cell, line and CHO-DG44 (B)/TNFR-Fc cell line, and protein-variety, GO enrichment and KEGG enrichment analysis were performed on the differential expression proteins. The results show that there are differences in cell growth, metabolism and exogenous protein expression between the two CHO cells. By comparing the protein expression profiles, it is found that there are 104 up-regulated proteins and 88 down-regulated proteins among 192 differentially expressed proteins. And enzymes accounted for the largest proportion (38.02%) among these differential proteins. The differentially expressed proteins are mainly enriched in biological processes like protein folding, glutathione metabolic process, carbohydrate metabolic process, intracellular protein transport, protein stabilization. These differential proteins are mainly related to pathways such as biosynthesis of amino acids, carbon metabolism, glycolysis/gluconeogenesis, citrate cycle, pyruvate metabolism and other signaling pathways. The differences in growth and metabolism, and expression of exogenous proteins between these two cell lines may be explained by the protomic differences in amino acid biosynthesis, carbon metabolism, glycolysis/gluconeogenesis, etc.
XIE Qiuling, TAN Chumin, WANG Yayu, et al . Proteomic Analysis of CHO-DG44 Cell Line Expressing Foreign Protein[J]. Journal of South China University of Technology(Natural Science), 2022 , 50(11) : 74 -81 . DOI: 10.12141/j.issn.1000-565X.220031
| 1 | LOUIE S, HEIDERSBACH A, BLANCO N,et al. Endothelial intercellular cell adhesion molecule 1 contri-butes to cell aggregate formation in CHO cells cultured in serum-free media [J].Biotechnology Progress,2020,36(3):e2951/1-14. |
| 2 | TIHANYI B, NYITRAY L .Recent advances in CHO cell line development for recombinant protein production [J].Drug Discovery Today:Technologies,2020,38:25-34. |
| 3 | REINHART D, DAMJANOVIC L, KAISERMAYER C,et al .Bioprocessing of recombinant CHO-K1,CHO-DG44,and CHO-S:CHO expression hosts favor either mAb production or biomass synthesis [J].Biotechnology Journal,2019,14(3):e1700686/1-11. |
| 4 | WURM M J, WURM F M .Naming CHO cells for bio-manufacturing:Genome plasticity and variant phenotypes of cell populations in bioreactors question the relevance of old names [J].Biotechnology Journal,2021,16(7):e2100165/1-13. |
| 5 | XU X, NAGARAJAN H, LEWIS N E,et al .The genomic sequence of the Chinese hamster ovary (CHO)-K1 cell line [J].Nature Biotechnology,2011,29(8):735-741. |
| 6 | BECKER J, TIMMERMANN C, JAKOBI T,et al .Next-generation sequencing of the CHO cell transcriptome [J].BMC Proceedings,2011,5():1-2. |
| 7 | KIM J Y, KIM Y G, LEE G M .CHO cells in biotechnology for production of recombinant proteins:current state and further potential [J].Applied Microbiology Biotechnology,2012,93(3):917-930. |
| 8 | ZHU G, SUN L, ALBANETTI T,et al .Quantitative analysis of the supernatant from host and transfected CHO cells using iTRAQ 8-plex technique [J].Biotechnology and Bioengineering,2016,113(10):2140-2148. |
| 9 | PASCOE D E, ARNOTT D, PAPOUTSAKIS E T,et al .Proteome analysis of antibody-producing CHO cell lines with different metabolic profiles [J].Biotechnology and Bioengineering,2007,98(2):391-410. |
| 10 | SCHABERT V F, WATSON C, JOSEPH G J,et al .Costs of tumor necrosis factor blockers per treated patient using real-world drug data in a managed care population [J].Journal of Managed Care Pharmacy,2013,19(8):621-630. |
| 11 | KUMAR A, BAYCIN-HIZAL D, WOLOZNY D,et al .Elucidation of the CHO Super-Ome (CHO-SO) by proteoinformatics [J].Journal of Proteome Research,2015,14(11):4687-4703. |
| 12 | YUK I H, NISHIHARA J, WALKER D,Jr.,et al .More similar than different:Host cell protein production using three null CHO cell lines [J].Biotechno-logy and Bioengineering,2015,112(10):2068-2083. |
| 13 | FISHER S, HANDRICK R, OTTE K .The art of CHO cell engineering:A comprehensive retrospect and future perspectives [J].Biotechnology Advances,2015,33(8):1878-1896. |
| 14 | LAKEN H A, LEONARD M W .Understanding and modulating apoptosis in industrial cell culture [J].Current Opinion in Biotechnology,2001,12(2):175-179. |
| 15 | 叶玲玲 .一种高通量定量检测细胞凋亡方法的建立和初步应用 [D].北京:中国人民解放军军事医学科学院,2004. |
| 16 | 肖政,范里,谭文松 .半乳糖流加对CHO细胞生长代谢及其表达的Fc融合蛋白糖基化的影响 [J].生物技术通报,2019,35(8):138-145. |
| 16 | XIAO Zheng, FAN Li, TAN Wen-song .Galactose feeding in CHO cell culture process:the impacts on the cell growth,metabolism and glycosylation of Fc fusion protein [J].Biotechnology Bulletin,2019,35(8):138-145. |
| 17 | SONG J, LEE K, PARK S W,et al .Lactic acid upregulates VEGF expression in macrophages and facilitates choroidal neovascularization [J].Investigative Ophthalmology & Visual Science,2018,59(8):3747-3754. |
| 18 | 高量,游磊,朱明龙,等 .rCHO细胞在葡萄糖限制流加培养过程中的生长与代谢特性 [J].高校化学工程学报,2006(1):74-78. |
| 18 | GAO Liang, YOU Lei, ZHU Ming-long,et al .Growth and metabolism of rCHO cells in glucose-limited fed batch culture [J].Journal of Chemical Engineering of Chinese Universities,2006(1):74-78. |
| 19 | JIANG H, MA N, SHANG Y,et al .Triosephosphate isomerase 1 suppresses growth,migration and invasion of hepatocellular carcinoma cells [J].Biochemical Biophysical Research Communications,2017,482(4):1048-1053. |
| 20 | REN X, ZOU Q, YUAN C,et al .The dominant role of oxygen in modulating the chemical evolution pathways of tyrosine in peptides:dityrosine or melanin [J].Angewandte Chemie International Edition,2019,58(18):5872-5876. |
| 21 | LI J, WONG C L, VIJAYASANKARAN N,et al .Feeding lactate for CHO cell culture processes:impact on culture metabolism and performance [J].Biotechno-logy and Bioengineering,2012,109(5):1173-1186. |
| 22 | GOLDRICK S, LEE K, SPENCER C,et al .On-Line control of glucose concentration in high-yielding mammalian cell cultures enabled through oxygen transfer rate measurements [J].Biotechnology Journal,2018,13(4):e1700607/1-10. |
/
| 〈 |
|
〉 |